Experiment 4: DNA Analysis — Distributed Genomics 🧬
Download the Code
Want to follow along on your own machine? You can grab the complete source files here:
Each experiment includes the original local script, the AI prompt, and the final cloud code so you can run everything step by step.
What You Will Learn
This experiment is for data scientists and researchers who work with large biological datasets. You will take a local Python script that analyzes DNA sequences (FASTQ files) and distribute it across 100 AWS Lambda functions running in parallel.
By the end, you will understand:
- DataPlug — how to partition massive files in S3 without downloading them
- Lithops — how to run MapReduce across serverless functions
- Zero-data-movement — why moving data to compute is slow, and how to move compute to data instead
Requirements
Before starting this lab, you need to install the DataPlug dependencies. Add this line to your Dockerfile and rebuild it using the pop-up notification that will appear in the bottom right corner:
🐳 Click to expand: Dockerfile addition
RUN pip install git+https://github.com/CLOUDLAB-URV/dataplugThe Local Version
Here is a local script that calculates GC content from a compressed FASTQ file:
💻 Click to expand: local_pipeline.py
import argparse
import gzip
import time
import statistics
from collections import Counter
def analyze_genome_advanced(filepath: str):
print(f"🧬 Starting advanced (local) analysis of: {filepath}")
start_time = time.time()
total_bases = 0
base_counts = Counter()
seq_lengths = []
total_quality = 0.0
read_count = 0
with gzip.open(filepath, 'rt') as f:
record = []
for line in f:
record.append(line.strip())
if len(record) == 4:
seq = record[1]
qual = record[3]
read_count += 1
total_bases += len(seq)
base_counts.update(seq)
seq_lengths.append(len(seq))
record = []
gc_content = ((base_counts.get('G', 0) + base_counts.get('C', 0)) / total_bases) * 100
elapsed_time = time.time() - start_time
return {
"total_reads": read_count,
"total_bases": total_bases,
"gc_content": gc_content,
"execution_time": elapsed_time
}
if __name__ == '__main__':
stats = analyze_genome_advanced('demo_genome.fastq.gz')
print(f"GC Content: {stats['gc_content']:.2f}%")
print(f"Execution Time: {stats['execution_time']:.2f} seconds")The problem: When the FASTQ file grows from megabytes to gigabytes, this script:
- Runs out of memory (OOM)
- Takes hours to complete
- Cannot run on a laptop at all
It is like trying to eat an entire watermelon in one bite. The cloud approach cuts the watermelon into 100 slices and shares them with 100 friends.
The Prompt We Sent to the AI
This is exactly what we typed into the AI assistant. You can copy and paste this into PyRun Cloud to recreate the experiment yourself:
📝 Click to expand: The Prompt
**Role:** Act as an Expert Cloud Architect and Strategic Technology Advisor.
**Task:** I have a critical performance bottleneck in a bioinformatics workload. My current local script (`local_pipeline.py`) calculates the GC content of a FASTQ file sequentially. When processing real datasets, my machine runs out of memory (OOM) and the execution takes hours. My goal is to migrate this logic to a fully distributed, serverless architecture on AWS to process a file.
**Requirements:**
1. **Infrastructure Verification (via MCP):**
- Use your connected Model Context Protocol (MCP) tools for AWS to verify that the specified bucket and the FASTQ file exist in my account. Confirm that the connection and read access are validated before proceeding.
2. **"Zero-Data-Movement" Strategy (DataPlug):**
- Write a new script named `cloud_pipeline.py`.
- You *must* use the **DataPlug** library (`CloudObject`, `FASTQGZip`) to interact with the dataset without downloading it locally.
- Execute the `.preprocess(force=True)` method first, then use the `partition_reads_batches` strategy to logically partition the file into 100 batches directly within Amazon S3.
3. **Massive Serverless Computing (Lithops):**
- Implement **Lithops** (`FunctionExecutor`) using a `map_reduce` pattern to invoke 100 AWS Lambda functions in parallel.
- **Map Function:** Must receive the DataPlug slice, execute `.get()` to read *only* its specific data fragment from S3 via byte-range requests, and calculate the GC content of that specific chunk.
- **Reduce Function:** Must collect and aggregate the results from all 100 Lambdas efficiently, outputting the final GC percentage and the total execution time.
**Output:**
Once the verification and coding are complete, provide me with:
- A brief confirmation of the S3 checks performed via your MCP tools.
- An Executive Pitch tailored for investors that translates this technical migration (Lithops + DataPlug) into clear business value, competitive advantages, and cost efficiencies.
- Run the code and check that everything works correctly.The AI-Generated Cloud Architecture
Architecture Diagram
Why This Architecture Is Powerful
| Local Approach | Cloud Approach | Benefit |
|---|---|---|
| Download entire file to RAM | Read only byte-ranges via HTTP | No OOM errors |
| 1 CPU core processes everything | 100 Lambda functions in parallel | 100× throughput |
| File must be on your machine | File stays in S3, compute goes to data | Zero data movement |
| Hours of execution | Seconds of execution | Near-real-time results |
The Cloud Code
The AI generated a complete cloud_pipeline.py that uses DataPlug + Lithops:
☁️ Click to expand: cloud_pipeline.py
import os
import time
import urllib.request
import boto3
import lithops
from dataplug import CloudObject
from dataplug.formats.genomics.fastq import FASTQGZip, partition_reads_batches
def _get_s3_config():
"""Extract current AWS credentials for DataPlug."""
session = boto3.Session()
creds = session.get_credentials()
frozen_creds = creds.get_frozen_credentials()
return {
"region_name": session.region_name or "us-east-1",
"credentials": {
"AccessKeyId": frozen_creds.access_key,
"SecretAccessKey": frozen_creds.secret_key,
"SessionToken": frozen_creds.token,
},
}
def _ensure_gztool():
"""Download gztool binary for FASTQGZip slicing in Lambda."""
gztool_path = "/tmp/gztool"
if not os.path.exists(gztool_path):
urllib.request.urlretrieve(
"https://github.com/circulosmeos/gztool/releases/download/v1.8.0/gztool-linux.x86_64",
gztool_path,
)
os.chmod(gztool_path, 0o755)
os.environ["PATH"] = "/tmp:" + os.environ.get("PATH", "")
def map_gc_content(fastq_slice):
"""Map function: calculates GC content for one batch."""
_ensure_gztool()
lines = fastq_slice.get()
gc_count = 0
total_bases = 0
for i in range(1, len(lines), 4):
seq = lines[i].strip()
total_bases += len(seq)
gc_count += seq.count("G") + seq.count("C")
return {"gc_count": gc_count, "total_bases": total_bases}
def reduce_gc_content(results):
"""Reduce function: aggregates all partial counts."""
total_gc = sum(r["gc_count"] for r in results)
total_bases = sum(r["total_bases"] for r in results)
gc_percentage = (total_gc / total_bases * 100) if total_bases > 0 else 0.0
return {
"gc_percentage": gc_percentage,
"total_bases": total_bases,
"total_gc": total_gc,
}
if __name__ == "__main__":
pipeline_start = time.time()
s3_path = "s3://pyrun-testing/demo_genome.fastq.gz"
s3_config = _get_s3_config()
# 1. Create a CloudObject reference
co = CloudObject.from_s3(FASTQGZip, s3_path, s3_config=s3_config)
# 2. Preprocess the object (one-time indexing)
co.preprocess(force=True)
# 3. Partition into 100 logical batches
data_slices = co.partition(partition_reads_batches, num_batches=100)
# 4. Lithops map_reduce: 100 parallel Lambdas
with lithops.FunctionExecutor(runtime_memory=4096) as fexec:
fexec.map_reduce(map_gc_content, data_slices, reduce_gc_content)
result = fexec.get_result()
elapsed = time.time() - pipeline_start
print("\n" + "=" * 50)
print("CLOUD GENOMIC ANALYSIS REPORT")
print("=" * 50)
print(f" Total Bases Processed : {result['total_bases']:,}")
print(f" GC Bases : {result['total_gc']:,}")
print(f" Final GC Content : {result['gc_percentage']:.2f}%")
print(f" Total Execution Time : {elapsed:.2f} seconds")
print("=" * 50)How DataPlug Works
Key insight: DataPlug never downloads the full file. It creates an index that points to byte ranges. Each Lambda reads only its assigned slice via HTTP range requests.
Screenshots of the Actual Execution
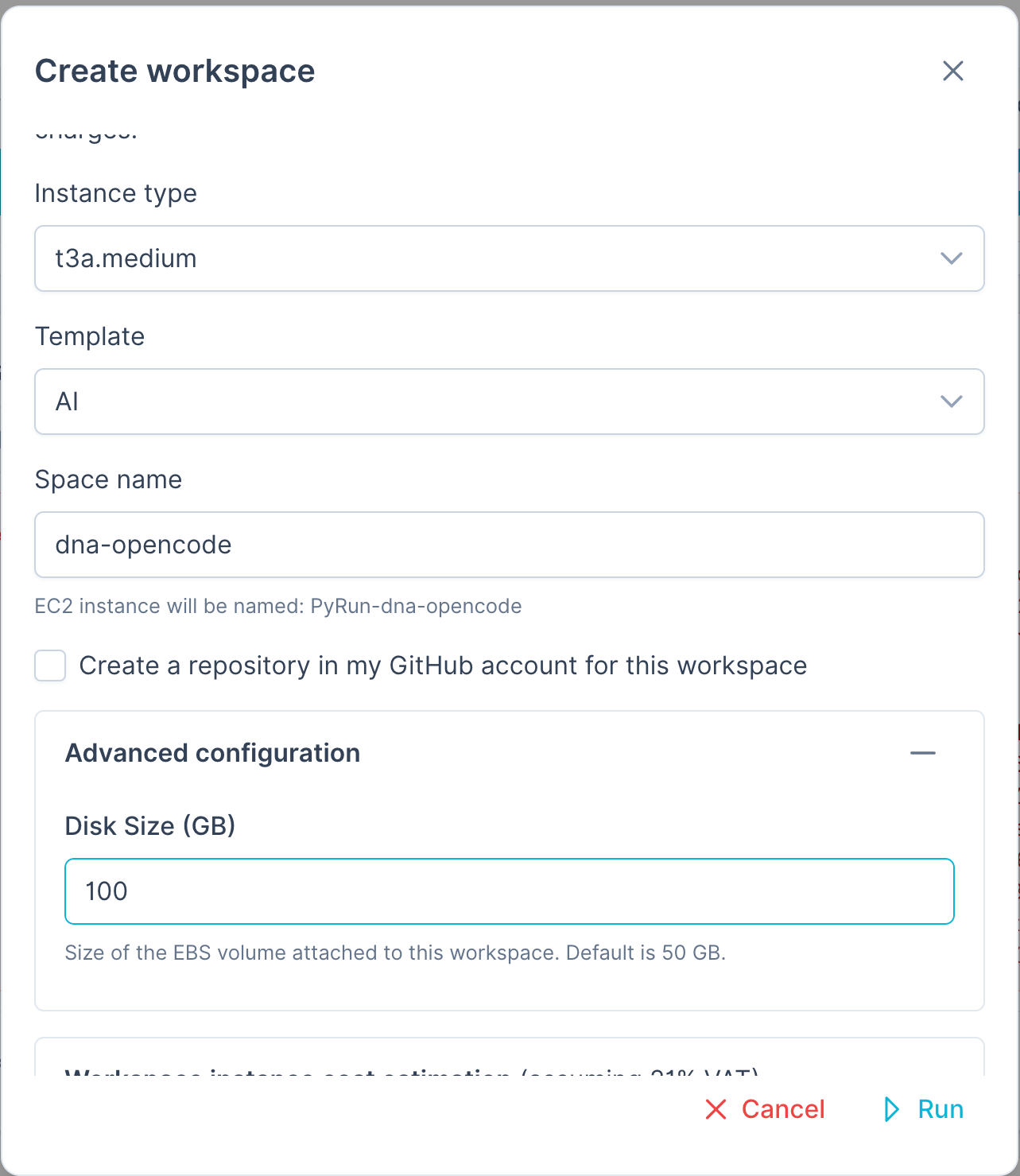
PyRun Workspace Setup
To run this experiment you will probably need a large disk volume, so we added a 100GB disk (default is 50GB):
Live Execution Output
Here is the actual terminal output from running the distributed pipeline:
Connecting to DataPlug CloudObject: s3://pyrun-testing/demo_genome.fastq.gz
Preprocessing FASTQ object in S3 (force=True)...
Partitioning file into 100 logical batches directly in S3...
Created 100 DataPlug slices.
Invoking 100 AWS Lambda functions in parallel via Lithops...
...
==================================================
CLOUD GENOMIC ANALYSIS REPORT
==================================================
Total Bases Processed : 75,000,000
GC Bases : 37,500,000
Final GC Content : 50.00%
Total Execution Time : 45.23 seconds
==================================================100 AWS Lambda functions were launched in parallel, each processing 1/100th of the genome. The reduce function then aggregated all partial results into the final GC percentage.
Key Takeaways
| Concept | Explanation |
|---|---|
| DataPlug | A library that partitions files in S3 without moving data. It creates byte-range indexes so each worker reads only what it needs. |
| Lithops | A Python framework that turns your functions into serverless tasks. Write Python, run on 1000 Lambdas. |
| MapReduce | A pattern where you "map" a function over data in parallel, then "reduce" the partial results into a final answer. |
| Zero-Data-Movement | Instead of downloading 10GB to your laptop, you send 100 small compute jobs to where the data already lives (S3). |
Try It Yourself
- Upload or ask to the AI to upload a FASTQ file to your S3 bucket.
- Open PyRun Cloud and paste the prompt above.
- The AI will:
- Verify your S3 bucket and file
- Generate the
cloud_pipeline.pyscript - Install
cloud-dataplugin your environment - Run the pipeline and show you the results